17 May 2021 by Remco Bouckaert
Understanding BEAST analyses
A BEAST analysis consists of a probabilistic model making up the posterior, operators, loggers, etc. A BEAST analysis can be drawn as a network like below
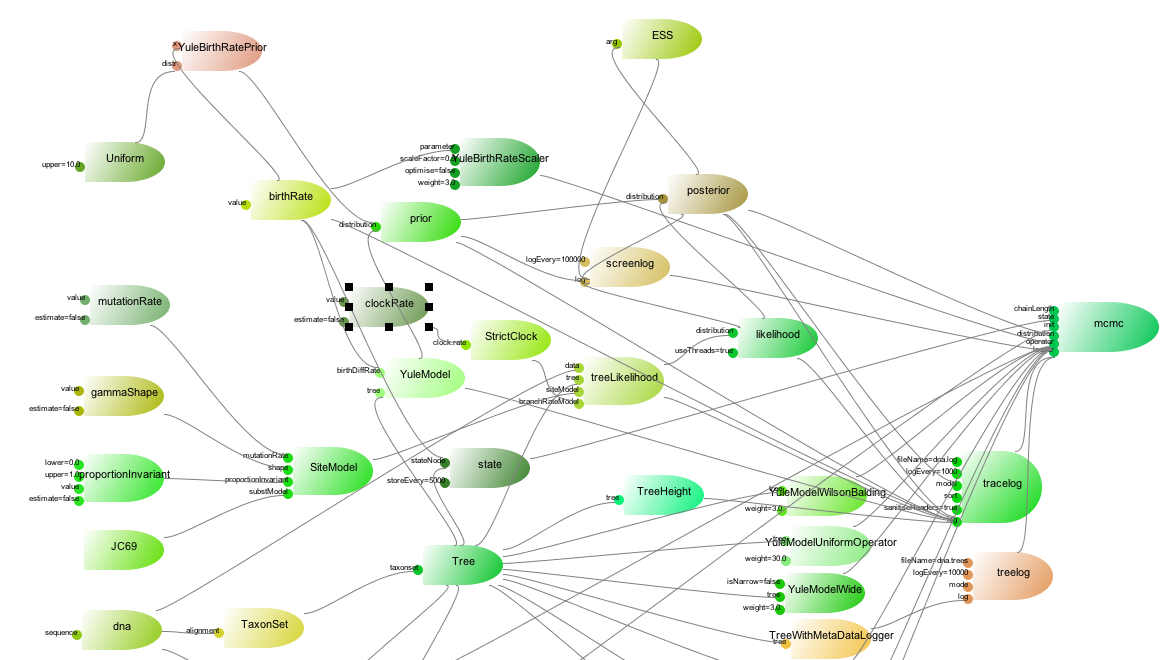
The ‘rockets’ represent BEAST objects, and the little thrusters inputs for each of the objects. Lines between rockets show how objects are connected to inputs. Note that cycles are not allowed: if A is connected to an input of B, B cannot be connected to an input of A, not even through a set of other objects C, D,….
For each rocket, there is a corresponding Java class, and the name of the class is specified in the spec attribute in the XML. The thrusters represent inputs to Java objects and are identified by names. Ther name attribute can be used to link objects to other objects (but more below).
Understanding XML
XML is a file format that is easy to read for computers, and arguably readable by humans as well. What we need to know for BEAST XML is that it consists of elements that start with < followed by the element name, e.g <element>. Every element is paired with a closing element that starts with </, e.g. </element>. Elements can be nested, like so <a><b></b></a>. If there are no elements inside an element, there is a shortcut for the end of element using the element name but ending with /> instead of >. So <a></a> can be written as <a/>.
So called attributes are properties that come with elements. Attributes are written as name="value" or if you prefer single quotes name='value'. Here is a small annotated example.
<element attribute="value" attribute2='value2'>
<!-- this is a comment -->
<inside></inside>
<insideWithShortCutForEnd/>
</element> <!-- end of element -->
As you can see, there are a number of special characters: <,>,",'. These cannot be used as attribute values: name="<-value" would be causing a parser error. To use these characters anyway, there is an escape character & followed by the name of the special character, and closed with semi-colon: < is <, > is >, " is ", ' is &;. But now we introduced another special character, & is &, so there are 5 of these ‘entities’. Other special characters not available in UTF-8 can be specified likewise.
Understanding BEAST XML
BEAST XML is based on having one element per BEAST object, as in the rocket-image above. Each thruster of a rocket represents an Input for a BEAST object that can be identified by a name. Therefore, you can create a BEAST analysis by specifying relations between BEAST objects and inputs of other BEAST objects. There are 4 reserved attribute names
spec= name of the BEAST class, for examplebeast.evolution.tree.RandomTree.name= name of the input of the object in the enclosing element.id= identifier of the object, useful for logging and referring form other places in the XML throughidrefs. The id should have a unique value.idref= reference to object elsewhere in the XML (not necessarily before, it can be specified later).
With this set of attributes, we can specify a random tree with constant population size of 1 like so:
<input id='RandomTree.t:dna' spec='beast.evolution.tree.RandomTree' name='init'>
<input name='estimate' value'false'/>
<input id='ConstantPopulation0.t:dna' name='populationModel'
spec='beast.evolution.tree.coalescent.ConstantPopulation' >
<input name='parameter' id='randomPopSize.t:dna' name='popSize'
spec='beast.core.parameter.RealParameter' value='1.0'/>
</input>
<input name='taxa' idref='dna'/>
<input name='initial' idref='Tree.t:dna'/>
</input>
Replace name attribute with element name
Note that we have not used the element name, which seems a bit of a waste. Instead of using any random element name (input in the above XML fragment), we can use the value of the name attribute. If the name attribute is not specified, BEAST assumes the element name value of the name attribute, and that this the input name to be connected to. So, we can write the above as:
<init id='RandomTree.t:dna' spec='beast.evolution.tree.RandomTree'>
<estimate value'false'/>
<populationModel id='ConstantPopulation0.t:dna'
spec='beast.evolution.tree.coalescent.ConstantPopulation'>
<parameter id='randomPopSize.t:dna' name='popSize'
spec='beast.core.parameter.RealParameter' value='1.0'/>
</populationModel>
<taxa idref='dna'/>
<initial idref='Tree.t:dna'/>
</init>
Moving simple elements inside attributes + @ notation for idrefs
Some primitive values can be marked up as attributes instead of specifying a separate element. For example, adding the estimate attribute to the init element and specifying its value (namely estimate="false") is interpreted as <estimate value'false'/>.
The idref construction occurs quite a lot in BEAST XML, and there is a shortcut in an attribute that uses @ to indicate we refer to an object and the value should not be taken literally. For example, taxa="@dna" is interpreted as <taxa idref='dna'/> and
initial="@Tree.t:dna" as <initial idref='Tree.t:dna'/>, so we get:
<init id="RandomTree.t:dna" taxa="@dna" initial="@Tree.t:dna" estimate="false"
spec="beast.evolution.tree.RandomTree" >
<populationModel id="ConstantPopulation0.t:dna"
spec="beast.evolution.tree.coalescent.ConstantPopulation">
<parameter id="randomPopSize.t:dna" name="popSize" value="1.0"
spec="beast.core.parameter.RealParameter"/>
</populationModel>
</init>
Note that this works with primitive values, but also with any BEAST class that has a String constructor. In particular, parameters like RealParameter have a String constructor, so instead of
<distr id="LogNormal1"
spec="beast.math.distributions.LogNormalDistributionModel">
<parameter id="popSizeLogM" estimate="false" name="M"
spec="beast.core.parameter.RealParameter" value="0"/>
<parameter id="popSizeLogS" estimate="false" name="S"
spec="beast.core.parameter.RealParameter" value="2"/>
</distr>
we can use
<distr id="LogNormal1" M="0.0" S="2.0"
spec="beast.math.distributions.LogNormalDistributionModel"/>
Namespace
To reduce the size of spec attributes, you can define a namespace in the beast element (the one at the first line of the XML). It looks like this by default:
namespace="beast.core:beast.evolution.alignment:beast.evolution.tree.coalescent:beast.core.util:beast.evolution.nuc:beast.evolution.operators:beast.evolution.sitemodel:beast.evolution.substitutionmodel:beast.evolution.likelihood"
When finding BEAST classes, first the spec attribute is used, but if it does not exist, it goes through the list of namespaces and appends the spec attribute to the namespace till it finds the class. So, instead of using spec="beast.evolution.tree.coalescent.ConstantPopulation" we can use spec="ConstantPopulation" and reduce the size of the XML a bit, so it looks like this:
<init id="RandomTree.t:dna" stimate="false" initial="@Tree.t:dna" taxa="@dna"
spec="beast.evolution.tree.RandomTree" >
<populationModel id="ConstantPopulation0.t:dna" spec="ConstantPopulation">
<parameter id="randomPopSize.t:dna" name="popSize" value="1.0"
spec="parameter.RealParameter"/>
</populationModel>
</init>
The plate element
BEAST recognises two special elements map and plate. Plates are useful when there is a lot of repetitive XML and are explained in the XML tricks post.
The value attribute
Placing any text inside an element is assigned to the value input. So, we have that
<parameter spec="parameter.RealParameter" name="popSize" value="1.0"/>
equals
<parameter spec="parameter.RealParameter" name="popSize">1.0</parameter>
The map element (discouraged)
The use of the map element is discouraged, but can be used to associated a spec attribute value with an element name. They should be specified inside the beast element, and outside the run element, for example with
<map name="LogNormal">beast.math.distributions.LogNormalDistributionModel</map>
any occurrence of a LogNormal element will be interpreted as <LogNormal spec="beast.math.distributions.LogNormalDistributionModel"..., for example:
<LogNormal id="LogNormal1" name="distr">
<parameter id="popSizeLogM" estimate="false" name="M"
spec="parameter.RealParameter">0</parameter>
<parameter id="popSizeLogS" estimate="false" name="S"
spec="parameter.RealParameter">2</parameter>
</LogNormal>
has no spec attribute on the LogNormal element, since the map already specified it.
XML errors
Mismatched open and end tags
[Fatal Error] :61:15: The element type "distribution" must be terminated by the matching
end-tag "</distribution>".
java.lang.IllegalArgumentException: org.xml.sax.SAXParseException; lineNumber: 61;
columnNumber: 15; The element type "distribution" must be terminated by the matching
end-tag "</distribution>"
This error can happen when some XML element is added and you accidentally forgot to put in a close tag somewhere. Note that the error message indicates an element, but the error may be much earlier in the XML.
In particular, it is possible that an element accidentally closed too early, like in the following line where the /> at the end should be >:
<prior id="YuleBirthRatePrior.t:dna" name="distribution" x="@birthRate.t:dna"/>
<Uniform id="Uniform.1" name="distr" upper="10"/>
</prior>
Just delete the / at the end of the first line to make this valid XML.
Duplicate attributes
[Fatal Error] :40:46: Attribute "spec" was already specified for element "run".
java.lang.IllegalArgumentException: org.xml.sax.SAXParseException; lineNumber: 40; columnNumber: 46; Attribute "spec" was already specified for element "run".
An attribute was already specified earlier, so just look up the element in the line number indicated and remove the attribute that is not wanted.
Missed close quote in an attribute
The intention was to have <log idref="posterior"/> but the end quote was forgotten, and <log idref="posterior/> was used. This gives the following error:
[Fatal Error] :106:10: The value of attribute "idref" associated with an element type "log" must not contain the '<' character.
java.lang.IllegalArgumentException: org.xml.sax.SAXParseException; lineNumber: 106; columnNumber: 10; The value of attribute "idref" associated with an element type "log" must not contain the '
Due to the missing close tag, the XML parser interprets the remaining XML as part of the attributes, and the <-character is not allowed in attribute values (use &lt; instead).
Missing end-of-element character
If the logger element is not properly closed, like so <logger id="screenlog" logEvery="100000" which misses the > at the end, we get this message:
[Fatal Error] :105:9: Element type "logger" must be followed by either attribute specifications, ">" or "/>".
java.lang.IllegalArgumentException: org.xml.sax.SAXParseException; lineNumber: 105; columnNumber: 9; Element type "logger" must be followed by either attribute specifications, ">" or "/>".
Extra character before opening beast element
The only part allowed before the beast element is the XML declaration (which contains some information about the XML format), which should look like this <?xml version="1.0" encoding="UTF-8" standalone="no"?>. If there is any character before or after that declaration, but before the opening beast element, you will get this error:
[Fatal Error] :1:55: Content is not allowed in prolog.
java.lang.IllegalArgumentException: org.xml.sax.SAXParseException; lineNumber: 1; columnNumber: 55; Content is not allowed in prolog.
Mismatched idref
It is easy to make a typo in an idref name, which results in this error:
Error 170 parsing the xml input file
Could not find object associated with idref posteriorX
Error detected about here:
<beast>
<run id='mcmc' spec='MCMC'>
<logger id='screenlog'>
<log>
Check the spelling of the attribute value of the idref. Sometimes, an extra @ snuck in, and <log idref="@posterior"/> gives a similar error, but with this messages:
Could not find object associated with idref @posterior
Likewise, if you forgot the @, for example in an operator that was supposed to look like <operator id="YuleBirthRateScaler.t:dna" spec="ScaleOperator" parameter="@birthRate.t:dna" scaleFactor="0.5" weight="3.0" optimise="false"/> but you accidentally put in parameter="birthRate.t:dna" without the @, the string birthRate.t:dna is interpreted as a parameter value, which results in this error:
Error 124 parsing the xml input file
Failed to set the string value to 'birthRate.t:dna' for beastobject id=YuleBirthRateScaler.t:dna
Error detected about here:
<beast>
<run id='mcmc' spec='MCMC'>
<operator id='YuleBirthRateScaler.t:dna' spec='ScaleOperator'>
Typo in spec attribute
When there is a typo in the spec attribute, the following error will be displayed:
Error 1017 parsing the xml input file
Class could not be found. Did you mean beast.core.MCMC?
Perhaps a package required for this class is not installed?
Error detected about here:
<beast>
<run id='mcmc' spec='MCMCM'>
In this case, an extra M ended up in spec="MCMCM", which should be spec="MCMC". However, it is also possible, that a package is missing. If you produced your XML with BEAUti, the opening beast element will have a required attribute that lists the packages used in the analysis, and an extra message appears before the above error message (in case package XYZ is not installed):
Error 1018 parsing the xml input file
Class MCMCM could not be found.
This XML requires the following package that is not installed: XYZ v1.0.0
See http://beast2.org/managing-packages/ for details on how to install packages.
Or perhaps there is a typo in spec and you meant beast.core.MCMC?
Error detected about here:
<beast>
<run id='mcmc' spec='MCMCM'>
